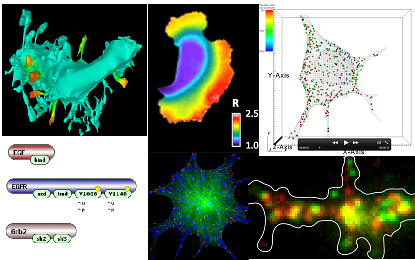
VCell is a unique computational environment for modeling and simulation of cell biology that is deployed as a distributed application freely available over the Internet. It is designed as a tool for a wide range of scientists, from experimental cell biologists to theoretical biophysicists. The creation of biological or mathematical models can range from the simple, to evaluate hypotheses or to interpret experimental data, to complex multi-layered models used to probe the predicted behavior of complex, highly non-linear systems. Models can be based on both experimental data and purely theoretical assumptions.
Users build complex models with a web-based Java interface to specify compartmental topology and geometry, molecular characteristics, and relevant interaction parameters. VCell automatically converts the biological description into a corresponding mathematical system of ordinary and/or partial differential equations. Distinct biological and mathematical frameworks are encompassed within a single graphical interface. The mathematics-savvy user may directly specify the complete mathematical description of the model, bypassing the schematic interface. VCell will then solve the equations by applying numerical solvers and generate appropriate software code to perform and analyze simulations. Results can be displayed and analyzed on-line or downloaded in a variety of formats.
Creating Models with VCell
The Virtual Cell Software has three primary documents for defining computational models different modeling approaches. Each document can be saved and loaded independently.
- BioModel: a graphical user interface environment to define model elements. A BioModel consists of a model description (Physiology), and one or more Applications that define conditions for exploring computational models of the Physiology.
- MathModel: a mathematical environment that specifies a mathematical model description either automatically generated from a BioModel Application or coded using the VCell Markup Language (VCML).
- Geometry: an explicitly defined 1D, 2D or 3D representation of a spatial structure that a BioModel Application or a MathModel can use for spatially resolved simulations.
The BioModel consists of:
- Physiology – a conceptual representation of the biological system in terms of cellular structures, species and biochemical reactions, including the types of kinetics that govern reactions. Rule-based modeling can also be used, which allows the representation of species as structured objects consisting of Molecules and uses reaction rules to define molecular interactions. The rule-based approach allows one to systematically incorporate site-specific details of molecular interactions into a model and simulate timecourses for model outputs (Observables) that may be functions of a large numbers of species.
- Pathway Information that associates elements of the model with annotated biological
- Applications that each define quantitative conditions needed to simulate and visualize a specific virtual experiment:
- Specifications for species initial conditions and reactions declarations, as well as network parameters in rule-based approaches
- Geometries used for simulations. Applications can be compartmental (well-mixed compartments) or can use different types of spatial geometries (see below).
- Protocols to define experimental events or electrical mapping.
- Parameter Estimation to estimate exact values of unknown parameters when the model is constrained by known datasets.
- A mathematical representation is automatically generated for each Application that can be viewed, or edited and run independently as a MathModel.
- Each Application can have multiple Simulations in which different numerical methods or conditions are used.
- Simulations with different solvers
- Simulations with different parameters (length of time, time step, spatial resolution, parameter overrides, etc.).
- Simulation Results can be viewed and analyzed in VCell or exported to a variety of formats.
The MathModel is a formalized math description of a BioModel Application. A declarative mathematical language describes processes modeled in the VCell. This language defines parameters, variables, and differential/algebraic systems defined over complex geometry.
- One or more Simulations must be defined for each MathModel to run and produce Results.
- The math from a BioModel Application is a valid MathModel, but a MathModel cannot be converted to a BioModel Application.
- MathModels can be used to override biology-driven limitations of BioModels.
The Geometry is a representation of a spatial structure that a BioModel Application (or a MathModel) can use for spatially resolved simulations.
- A geometry can be 1-D, 2-D, or 3-D.
- Geometry can be defined in multiple ways:
- Analytically, using mathematical equations
- Using provided templates for basic shapes
- Based on an imported image set
- Created using solid geometry (translate-rotate-scale)
VCell Capabilities
Model specification
- User convenience features:
- Local and global parameters
- Initial conditions can be specified in terms of other model parameters, reserved symbols and species.
- Autocomplete for parameter and species names
- Search and sorting for better viewing of large models
- Concentrations, surface densities and/or molecule counts for species
- Annotations for all model elements include text and links to various public databases
- Spatial features:
- Compartmental volumes and membrane sizes
- 3D Diffusion
- 2D Membrane Diffusion
- Advection (Velocity)
- 3D surface visualization for geometry
- Field Data to use experimental image data sets to define initial conditions for spatial simulations or to use the output of one spatial simulation as initial condition for another simulation.
- Rule-based specifications:
- Molecules – structured objects with binding sites having optional attribute states, such as phosphorylation
- Species are made of molecules (optionally) connected through binding sites.
- Reaction rules describe transformations of species classes by specifying the initial and final states of molecules participating in the reaction.
- Observables that describe classes of species with specified features, e.g. having the same site in a phosphorylated state.
Simulation capabilities
- Non-Spatial Deterministic (ODE) Solvers
- Forward Euler (First Order, Fixed Time Step)
- Runge-Kutta (Second Order, Fixed Time Step)
- Runge-Kutta (Fourth Order, Fixed Time Step)
- Adams-Moulton (Fifth Order, Fixed Time Step)
- Runge-Kutta-Fehlberg (Fifth Order, Variable Time Step)
- IDA (Variable Order, Variable Time Step, ODE/DAE)
- CVODE (Variable Order, Variable Time Step)
- Combined stiff solver CVODE/IDA
- Spatial Deterministic (PDE) Solvers
- Semi-Implicit Finite Volume, Regular Grid (Fixed Time Step)
- Semi-Implicit Finite Volume Compiled, Regular Grid (Fixed Time Step)
- Fully-Implicit Finite Volume, Regular Grid (Fixed Time Step).
- Non-Spatial Stochastic Solvers
- Gibson (Next Reaction Stochastic Method)
- Hybrid (Gibson + Euler-Matuyama Method)
- Hybrid (Gibson + Milstein Method)
- Hybrid (Adaptive Gibson + Milstein Method)
- Spatial Stochastic Solvers
- Smoldyn (“Smoluchowski Dynamics” written by Dr. Steve Andrews)
- Network-Free Stochastic solver
- NFSim (“Network-Free Simulator” written by Sneddon, Faeder & Emonet)
Parameter Estimation using COPASI for Non-Spatial Deterministic Problems
The Virtual Cell incorporates the COPASI parameter estimation capabilities to optimize parameters in non-spatial deterministic models to best fit experimental data. The available optimization solvers are listed below:
- Evolutionary Programming
- Evolution Strategy (SRES)
- Genetic Algorithm
- Genetic Algorithm SR
- Hooke and Jeeves
- Levenberg – Marquardt
- Nelder – Mead
- Particle Swarm
- Random Search
- Simulated Annealing
- Steepest Descent
- Praxis
- Truncated Newton
VCell Import
- SBML, SED-ML and BNGL formats
- Annotated SBML models from the BioModels database
- Pathway information from Pathway Commons
VCell Export
- Native VCML
- SBML L3 Core and Spatial
- SED-ML and COMBINE archive (.omex)
- Rule-based models in BNGL and NFSim XML
- MatLab m-files for systems of ODE’s
- formal pdf report
- Reaction diagram screenshot
- Simulation results in HDF5 format
Simulation results
- Non-Spatial Deterministic Simulations
- Plots and tabular data for species, functions, observables, and reaction fluxes
- Non-Spatial Stochastic Simulations
- Plots and tabular data for species, functions, observables, and reaction fluxes
- Representation as concentration and molecule counts
- Single, multiple trajectories or a histogram of values at a given time point.
- Spatial Deterministic Simulations:
- Timecourses and kymographs over specified regions of interest (ROI) – points, lines, curves, areas
- Predefined ROIs for existing spatial compartments.
- Post-processing of statistical data over ROI
- Various methods to define ROIs:
- Analytic equations
- User defined Boolean functions.
- User drawn polygons.
- Save results as
- Raw data
- Images (Tiff, JPEG, GIF)
- Movies
- VTK unstructured format (.vtu) for the Visualization Toolkit (http://vtk.org),
- Nearly Raw Raster Data (.nrdd)
VCell Team
VCell Administrators
Principal Investigator
Leslie M. Loew, Ph.D.
UConn Board of Trustees Distinguished Professor
Boehringer Ingelheim Chair in Cell Sciences
Professor, Department of Cell BiologyCenter for Cell Analysis and Modeling
Email: les@uchc.edu
Phone: 860-679-3568
Systems Administrator
Ion I. Moraru, M.D., Ph.D.
Professor, Department of Cell Biology
Center for Cell Analysis and Modeling
Email: moraru@uchc.edu
Phone: 860-679-2908
Numerical Methods Lead
Boris Slepchenko, Ph.D.
Associate Professor, Department of Cell Biology
Center for Cell Analysis and Modeling
Email: boris@uchc.edu
Phone: 860-679-4025
Associated Personnel
Alumni
Contact
Center for Cell Analysis and Modeling
UConn Health
263 Farmington Avenue
Farmington, CT 06030-6406
| VCell Director | Administrative Contact |
|---|---|
| Les Loew | Karen Zucker |
| les@volt.uchc.edu | zucker@uchc.edu |
| Phone: 860-679-3568 | Phone: 860-679-1452 |
| Fax: 860-679-1039 |